Renowned Hungarian physicist Albert-László Barabási and his colleagues have reached another milestone in their coronavirus research. The group has published its findings on the effectiveness of drugs approved for other diseases for SARS-CoV-2 infections. They used different network repurposing methodologies to test a wide range of compounds. This new approach can significantly accelerate the process of coronavirus drug development.
In their latest study, “Network Medicine Framework for Identifying Drug Repurposing Opportunities for COVID-19“, Albert-László Barabási and his group identified numerous potential drug candidates for COVID-19 and proposed an algorithmic toolset to identify future treatments for diseases underserved by conventional drug discovery processes, Portfolio reported. The Hungarian-American physicist, best known for his work in the research of network theory, announced the great news on his Facebook page:
By applying three network repurposing methodologies (artificial intelligence-based algorithm, diffusion algorithm, and proximity algorithm), the research group examined approximately 6,000 FDA-Approved (Food and Drug Administration) Drugs for their expected efficacy against SARS-CoV-2. The first list of 918 drugs for which all pipelines offered predictions was published in April. The group pre-incubated the Vero cells with these drugs and tested their effect on the virus.
“Of the 918 drugs, 806 had no detectable effect on viral infectivity (N drugs, 87.8% of the tested list); 35 were cytotoxic to the host cells (C drugs); 37 had a strong effect (S drugs), being active over a broad range of concentrations; and 40 had a weak effect (W drugs) on the virus.”
The researchers have found 77 drugs with a positive outcome (S&W) from in vitro screen.
After further inspection, they revealed that only one of the 77 S&W drugs targets a viral protein binding directly. 76 can be considered “network drugs” which cannot be identified using traditional binding-based methods because they achieve their effect indirectly by perturbing the host subcellular network.
The authors argued that “no single method offers consistently reliable outcomes across all datasets and metrics. This prompted us to
develop a multimodal approach that fuses the predictions of all algorithms, showing that a consensus among the different predictive methods consistently exceeds the performance of the best individual pipelines.”
The rapid spread of the coronavirus pandemic created a growing need for methodologies that can quickly and reliably identify clinically approved compounds that may be effective in the treatment of coronavirus patients. Drug repurposing algorithms rank drugs according to various criteria, allowing for the time-effective analysis of a wide range of compounds.
The researchers added that
“the lack of reliable repurposing methodologies has resulted in a winner-takes-all pattern, where more than one-third of registered clinical trials focus on hydroxychloroquine or chloroquine, siphoning away resources from testing a wider range of potentially effective drug candidates.”
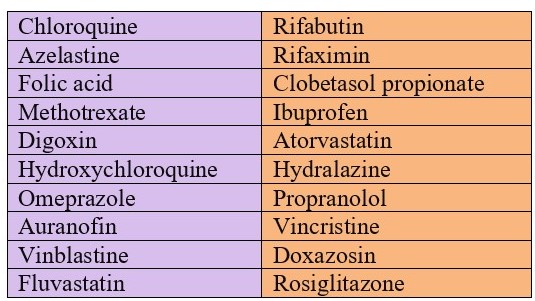
Top 10 drugs with strong and weak effect on COVID-19: